pacman::p_load(tmap, tidyverse, sf)In-class Exercise 3: Analytical Mapping
In-class Exercise 3: Analytical Mapping
Getting Started
Loading Packages
Importing Data
NGA_wp <- read_rds("data/rds/NGA_wp.rds")Choropleth Mapping
Distribution of non-functional water point
Plot a choropleth map showing the distribution of non-function water point by LGA
p1 <- tm_shape(NGA_wp) +
tm_fill("wp_functional",
n = 10,
style = "equal",
palette = "Blues") +
tm_borders(lwd = 0.1,
alpha = 1) +
tm_layout(main.title = "Distribution of functional water point by LGAs",
legend.outside = FALSE)p2 <- tm_shape(NGA_wp) +
tm_fill("total_wp",
n = 10,
style = "equal",
palette = "Blues") +
tm_borders(lwd = 0.1,
alpha = 1) +
tm_layout(main.title = "Distribution of total water point by LGAs",
legend.outside = FALSE)tmap_arrange(p2, p1, nrow = 1)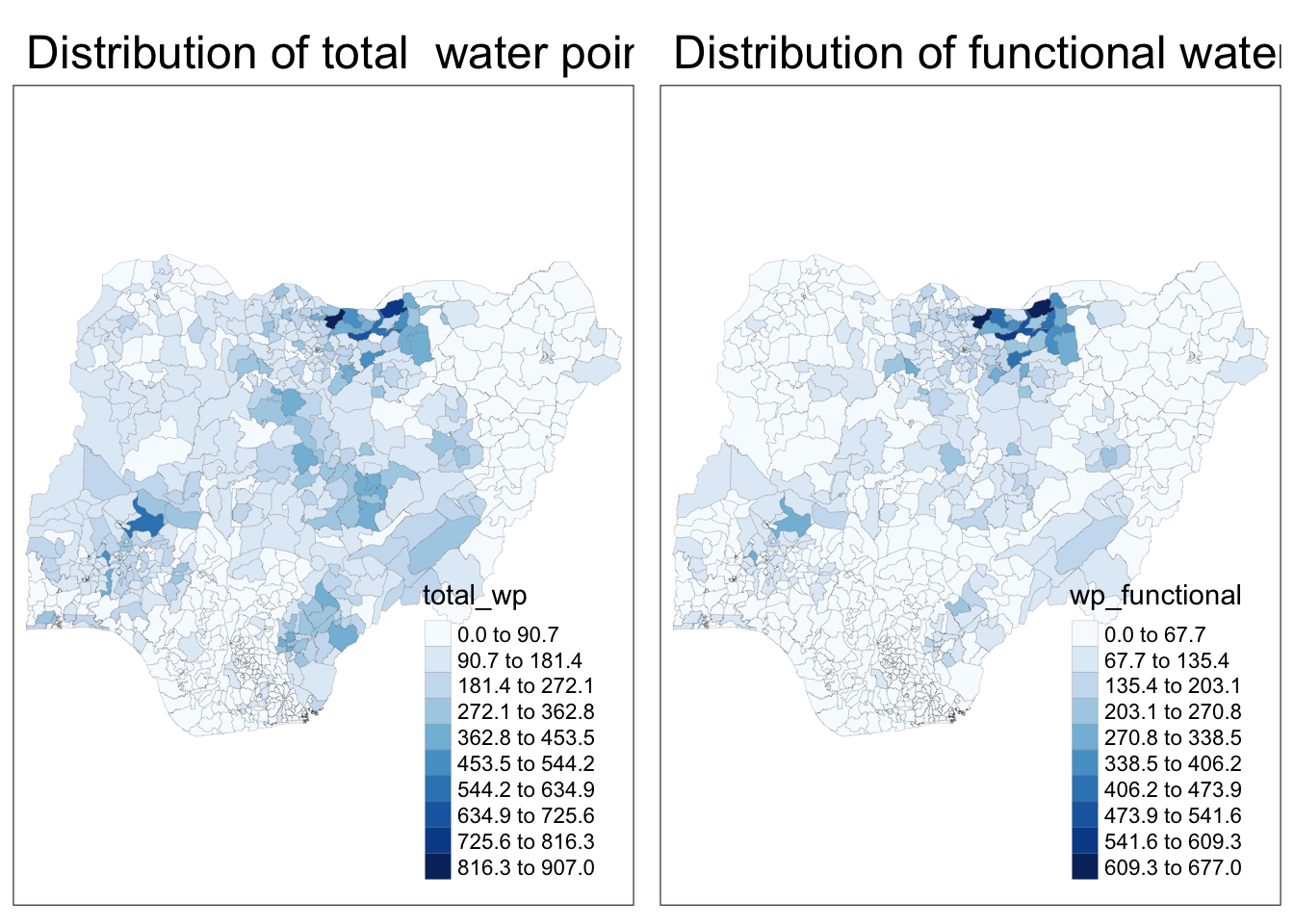
Choropleth Map for Rates
Deriving Proportion of Functional Water Points and Non-Functional Water Points
NGA_wp <- NGA_wp %>%
mutate(pct_functional = wp_functional/total_wp) %>%
mutate(pct_nonfunctional = wp_nonfunctional/total_wp)Plotting map of rate
Plot a choropleth map showing the distribution of percentage functional water point by LGA
tm_shape(NGA_wp) +
tm_fill("pct_functional",
n = 10,
style = "equal",
palette = "Blues",
legend.hist = TRUE) +
tm_borders(lwd = 0.1,
alpha = 1) +
tm_layout(main.title = "Rate map of functional water point by LGAs",
legend.outside = TRUE)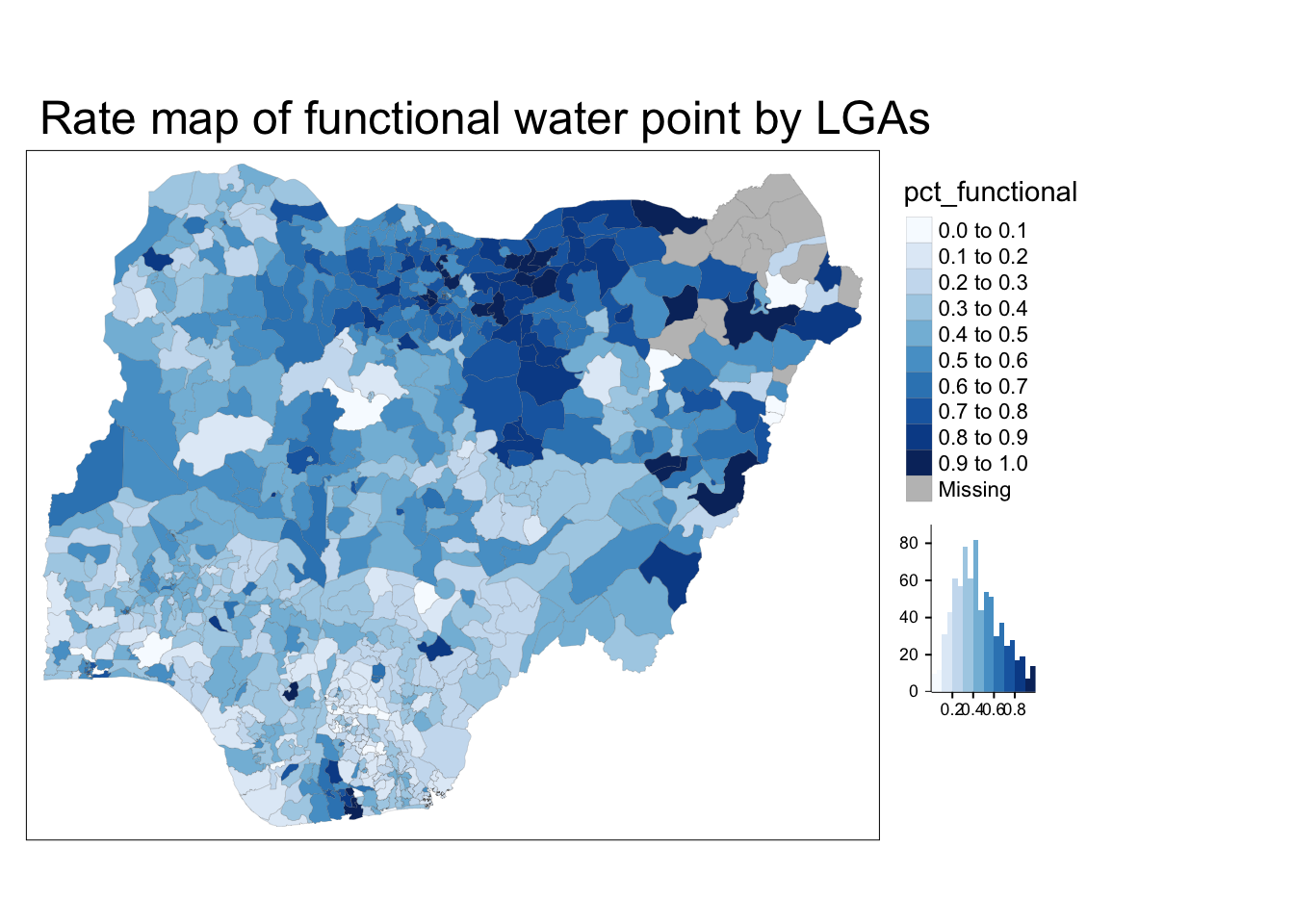
Extreme Value Maps
Percentile Map
Data Preparation
#Exclude records with NA by using the code chunk below
NGA_wp <- NGA_wp %>%
drop_na()#Creating customised classification and extracting values
percent <- c(0,.01,.1,.5,.9,.99,1)
var <- NGA_wp["pct_functional"] %>%
st_set_geometry(NULL)
quantile(var[,1], percent) 0% 1% 10% 50% 90% 99% 100%
0.00000000 0.01818182 0.18181818 0.41666667 0.76086957 1.00000000 1.00000000
Important
When variables are extracted from an sf data.frame, the geometry is extracted as well. For mapping and spatial manipulation, this is the expected behavior, but many base R functions cannot deal with the geometry. Specifically, the quantile() gives an error. As a result st_set_geomtry(NULL) is used to drop geomtry field.
Creating get.var function
get.var <- function(vname,df){
v <- df[vname]%>%
st_set_geometry(NULL)
v <- unname(v[,1])
return(v)
}Percentile Mapping Function
percentmap <- function(vnam, df, legtitle=NA, mtitle="Percentile Map"){
percent <- c(0,.01,.1,.5,.9,.99,1)
var <- get.var(vnam, df)
bperc <- quantile(var, percent)
tm_shape(df) +
tm_polygons() +
tm_shape(df) +
tm_fill(vnam,
title=legtitle,
breaks=bperc,
palette="Blues",
labels=c("< 1%", "1% - 10%", "10% - 50%", "50% - 90%", "90% - 99%", "> 99%")) +
tm_borders() +
tm_layout(main.title = mtitle,
title.position = c("right","bottom"))
}Percentile Map
percentmap("total_wp", NGA_wp)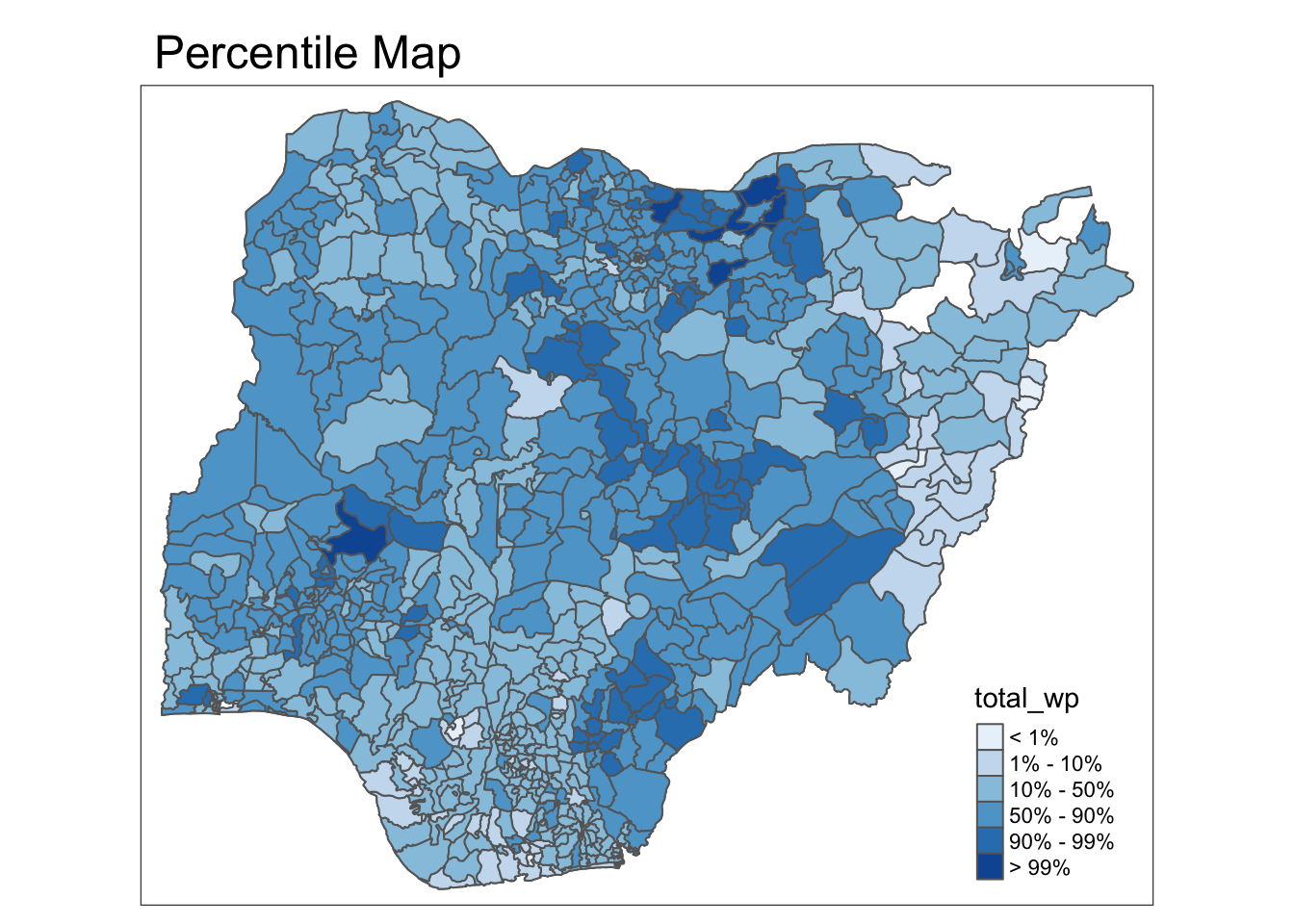
Box Map
ggplot(data = NGA_wp,
aes(x = "",
y = wp_nonfunctional)) +
geom_boxplot()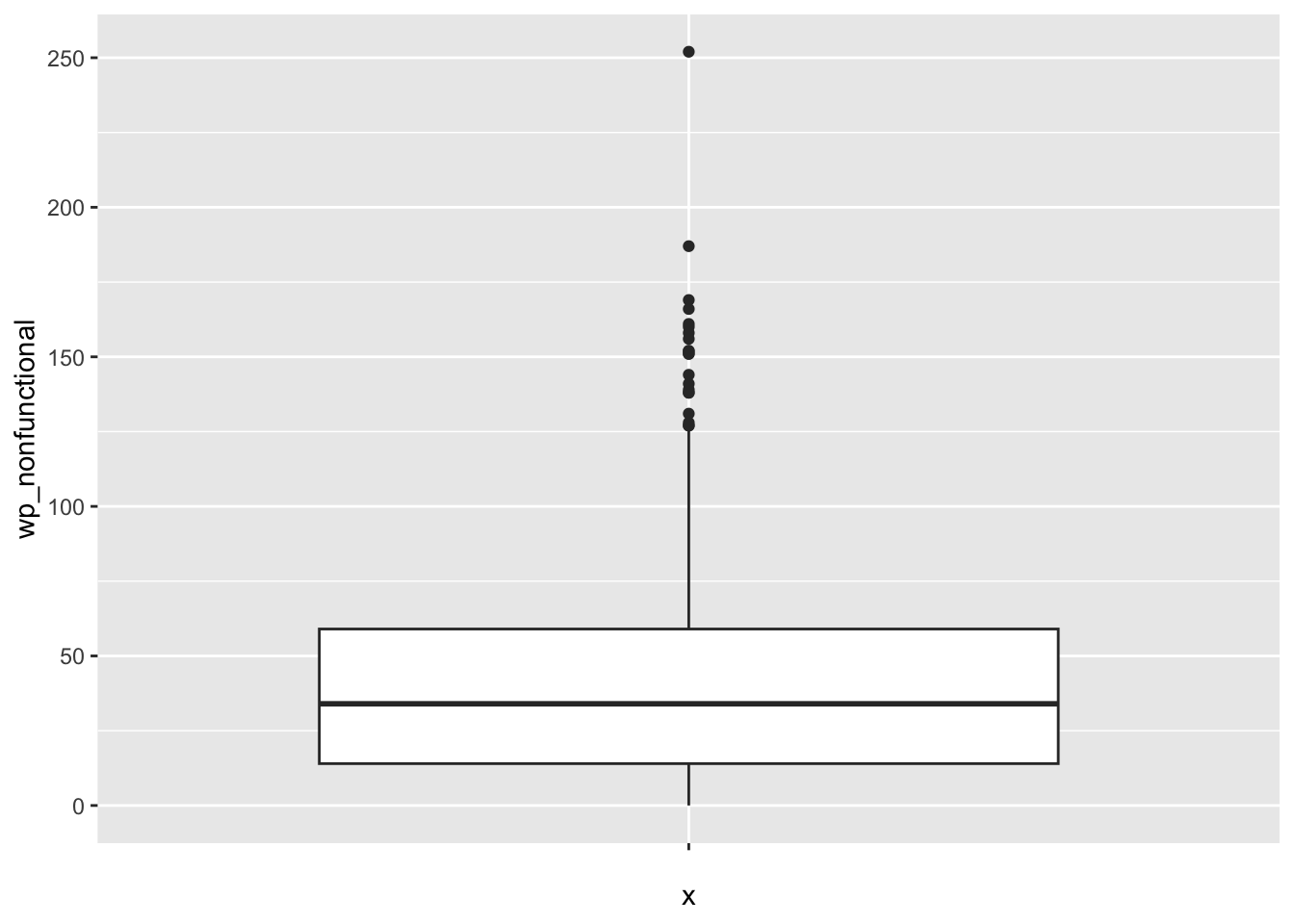
Creating boxbreaks function
boxbreaks <- function(v,mult=1.5) {
qv <- unname(quantile(v))
iqr <- qv[4] - qv[2]
upfence <- qv[4] + mult * iqr
lofence <- qv[2] - mult * iqr
# initialize break points vector
bb <- vector(mode="numeric",length=7)
# logic for lower and upper fences
if (lofence < qv[1]) { # no lower outliers
bb[1] <- lofence
bb[2] <- floor(qv[1])
} else {
bb[2] <- lofence
bb[1] <- qv[1]
}
if (upfence > qv[5]) { # no upper outliers
bb[7] <- upfence
bb[6] <- ceiling(qv[5])
} else {
bb[6] <- upfence
bb[7] <- qv[5]
}
bb[3:5] <- qv[2:4]
return(bb)
}Creating get.var function
get.var <- function(vname,df) {
v <- df[vname] %>% st_set_geometry(NULL)
v <- unname(v[,1])
return(v)
}Testing
var <- get.var("wp_nonfunctional", NGA_wp)
boxbreaks(var)[1] -53.5 0.0 14.0 34.0 59.0 126.5 252.0Boxmap Function
boxmap <- function(vnam, df,
legtitle=NA,
mtitle="Box Map",
mult=1.5){
var <- get.var(vnam,df)
bb <- boxbreaks(var)
tm_shape(df) +
tm_polygons() +
tm_shape(df) +
tm_fill(vnam,title=legtitle,
breaks=bb,
palette="Blues",
labels = c("lower outlier",
"< 25%",
"25% - 50%",
"50% - 75%",
"> 75%",
"upper outlier")) +
tm_borders() +
tm_layout(main.title = mtitle,
title.position = c("left",
"top"))
}Box Map
tmap_mode("plot")
boxmap("wp_nonfunctional", NGA_wp)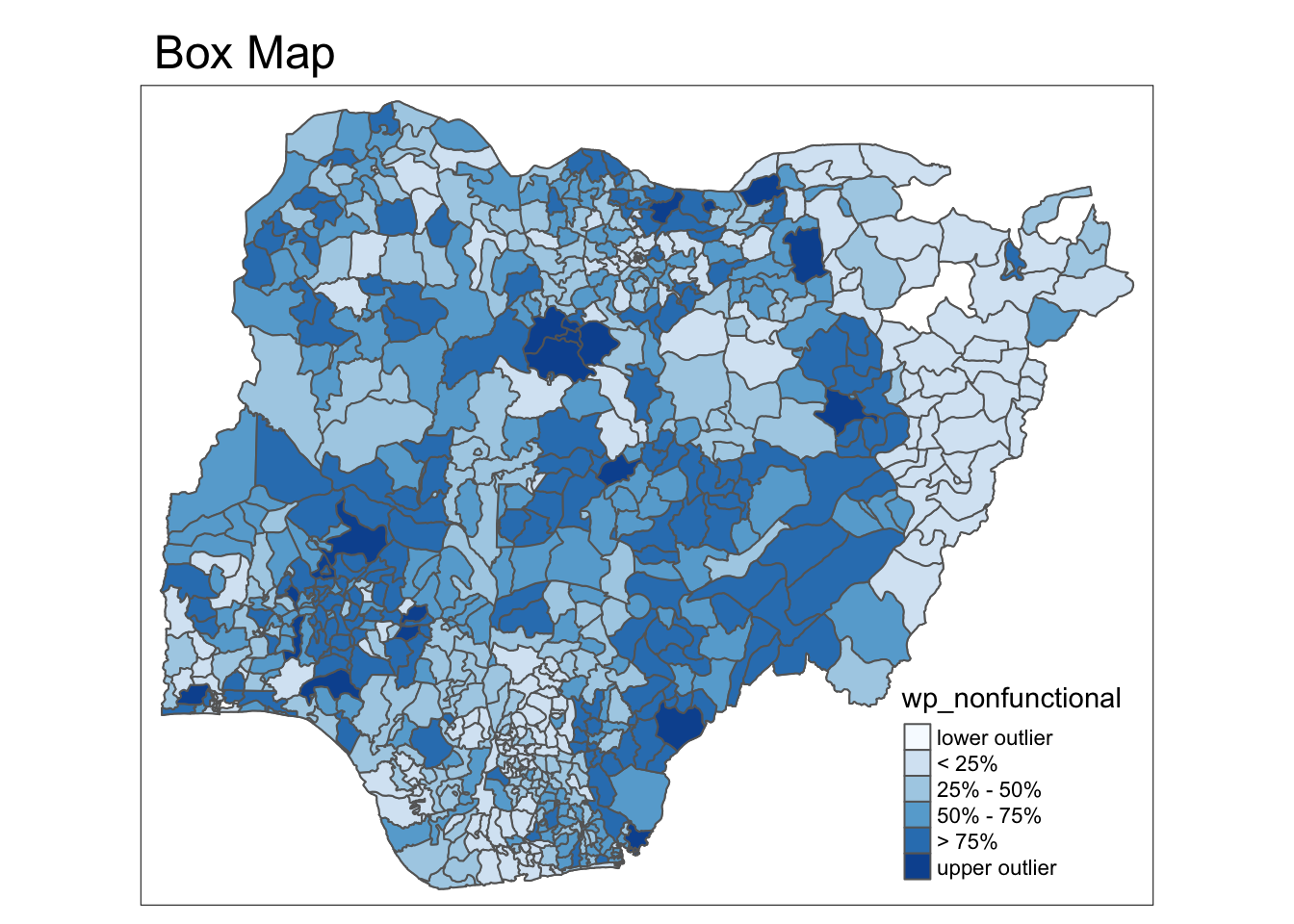
Recode zero
The code chunk below is used to recode LGAs with zero total water point into NA.
NGA_wp <- NGA_wp %>%
mutate(wp_functional = na_if(
total_wp, total_wp < 0))